BMP signalling: development, stem cell maintenance and cancer
The Bone Morphogenic Proteins (BMPs) represent a major, highly conserved family of growth factor signalling molecules. The BMP signalling pathway is a key regulator of a diverse array of processes, including the patterning of many organs and tissues during embryonic development. Consequently, misregulation of BMP signalling is associated with a spectrum of human diseases, including different cancer types, vascular disorders, kidney diseases and skeletal defects.
The Ashe lab uses the model organism Drosophila melanogaster to investigate the regulation of BMP signalling. In Drosophila, the major BMP signalling molecule is called Decapentaplegic (Dpp), and is important during numerous stages of development. The primary aim of this group is to understand how Dpp signalling is regulated and interpreted in the early embryo and the ovary. In the ovary Dpp is a critical signal required to maintain germline stem cells in the niche. In the embryo a gradient of Dpp activity leads to the specification of different cell types.
Past projects in this lab have been based around transcription regulation and dynamics of Dpp target genes in the early embryo, the extracellular regulation of the Dpp pathway and the maintenance of Germline Stem Cells (GSCs) in the germarium.
Gene Expression Dynamics during Embryogenesis
Transcription Kinetics
To investigate BMP gradient interpretation at single-cell resolution in the early embryo we have recently utilised a combination of the MS2/MCP live imaging system and computational modelling. MS2 stem-loops were inserted in the endogenous 5’UTR of BMP target genes hindsight (hnt) and u-shaped (ush) using CRISPR/Cas-9 genome engineering. The stem-loops are bound specifically by the MS2 coat protein (MCP), which is fused to GFP and allows us to visualise nascent transcription. A memory-adjusted Hidden Markov Model was used to infer the promoter state and burst parameters of nuclei in different regions of the BMP gradient at single-cell resolution. This study revealed that BMP signalling modulates burst frequency by regulating the promoter activation rate (Hoppe et al 2020, Developmental Cell).
The MS2/MCP Live Imaging System
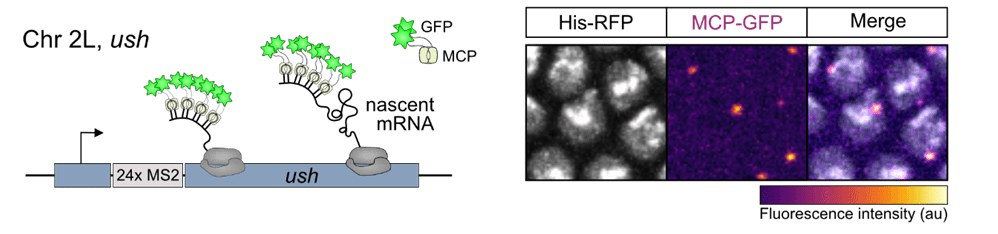
Transcription Regulation
Using a combined approach of RNA-seq with Mad and Brinker ChIP-seq in the early embryo we identified multiple BMP target genes and their enhancers, including regulators of EGF signalling which we showed to negatively regulate BMP-responsive transcription. We have also investigated how motifs in the BMP-responsive enhancers contribute to expression patterns, and demonstrated a key role for the pioneer factor Zelda in BMP gradient interpretation (Deignan et al 2016, PLoS Genetics). Additionally, we have used Global Run On sequencing (Gro-seq) to map RNA polymerase density during early embryogenesis and have identified widespread promoter proximal pausing in BMP target genes and pathway components (Saunders et al 2013, Genes and Development).
Extracellular Regulation of BMPs
BMP Shuttling Model
In the Drosophila embryo, BMP gradient formation requires shuttling of a heterodimer of BMPs Dpp and Screw (Scw) to the dorsal midline. We previously found that type IV collagens, encoded by the vkg and Dcg1 genes in Drosophila, bind Dpp and are important for Dpp gradient formation in the early embryo (Wang et al, 2008, Nature). We have mapped additional protein interactions allowing us to formulate a molecular multistep model of Dpp shuttling complex assembly on collagen IV (Sawala et al, 2012, PNAS).
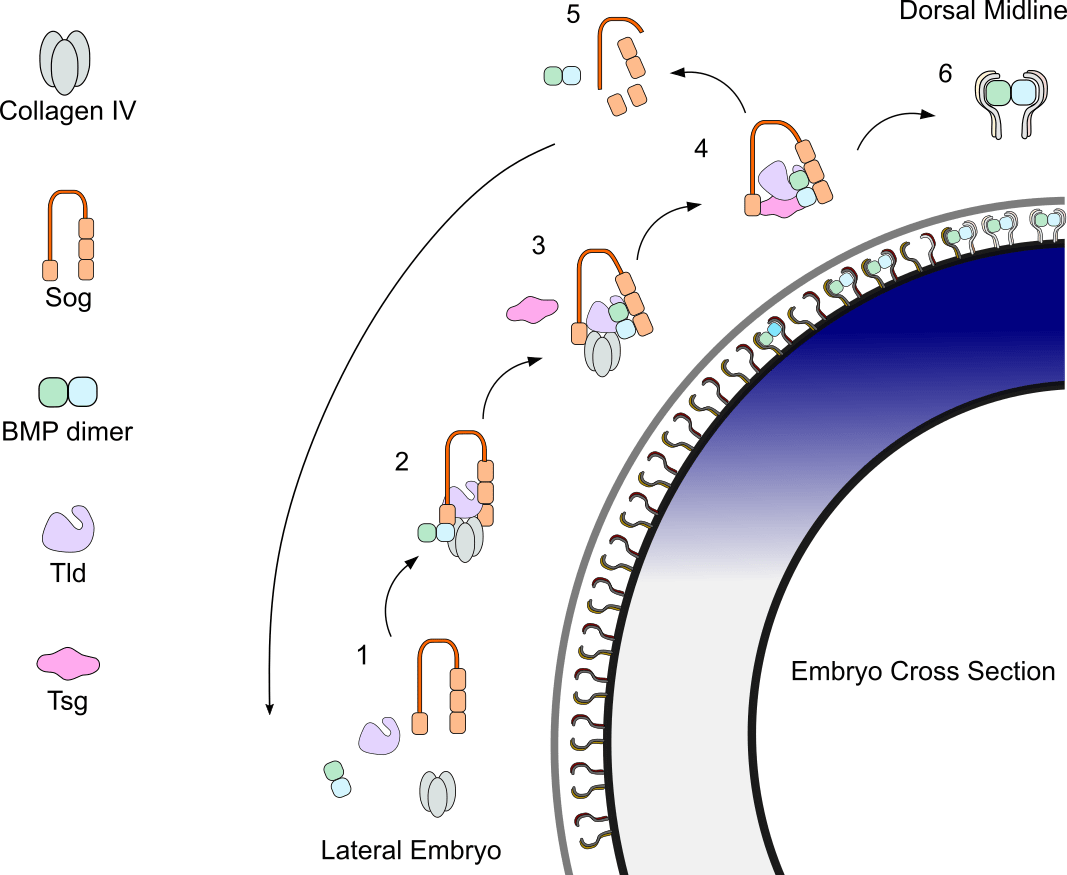


Shuttling complex assembly and release from Collagen IV to achieve peak BMP signalling
-
- BMP heterodimer levels are highest in dorsal regions. Sog levels are highest in ventral regions.
- An inhibitory complex is formed in dorsolateral regions; the BMP dimer is bound by the extracellular antagonist Sog and metalloproteinase Tolloid, forming a complex with the scaffold protein Collagen IV.
- Shuttling complex is released from Collagen IV through Tsg binding.
- Shuttling complex is free to diffuse dorsally.
- Sog is cleaved by Tolloid. BMP dimer is released. Due to low levels of Sog dorsally the complex is not rebound.
- BMP dimer binds receptor
Subsequently, we identified a requirement for integrins downstream of Collagen IV in peak BMP signalling. Integrins amplify the BMP signal by forming complexes with BMP receptors, notably through a direct interaction between BMP receptor Thickveins and the cytoplasmic tail of integrin βPS, and enhancing Mad phosphorylation. (Sawala et al 2015, Cell Reports).
Structural Analysis
In collaboration with Clair Baldock’s lab we undertook a structural analysis of the metalloprotease Tolloid (Tld). Using Small Angle X-ray Scattering (SAXS) and transmission electron microscopy we showed that the two C-terminal CUB domains of Tld mediate Short gastrulation (Sog) interaction. Additionally, Tld interacts with Collagen IV via its N-terminal CUB domain, which facilitates Sog cleavage. We also recapitulated embryonic BMP gradient formation synthetically in the absence of Tld’s non-catalytic domains, by using leucine zippers to artificially tether the Tld protease domain to Sog (Winstanley et al, 2015, eLife).
Germline Stem Cell Maintenance in the Germarium
Cytosensor Projections
We have used RNA-seq to identify the transcriptomes in the Germline Stem Cells (GSCs) and Cystoblasts (CBs), as well as Dpp-target genes in GSCs. The GSC transcriptome has provided new information about how GSCs respond to the Dpp signal. We identified two different types of cellular projection produced by the GSCs which function to receive the niche Dpp signal; short actin-based projections and longer actin and microtubule rich projections, which we call ‘cytosensors’. Both types of projection enable GSCs to access a Dpp reservoir located away from the niche GSC interface. This Dpp reservoir is maintained by the glypican Dally sequestering Dpp at the anterior face of niche cells. Cytocensors, which form downstream of Dpp, have an additional function in attenuating the Dpp signal, enabling fine tuning of the signalling level in the GSC (Wilcockson & Ashe, 2019, Developmental Cell).
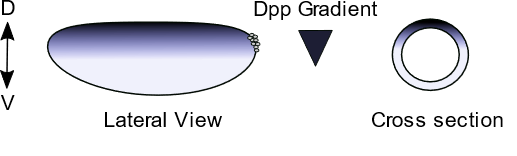
Drosophila Germarium

The GSC synthesises multiple actin filopodia to access a Dpp reservoir (right panel) localised at the anterior of the niche (top projection, one is shown for simplicity). A microtubule (green) and actin (red) based cytocensor projection (lower projection) forms downstream of Dpp to collect and attenuate Dpp signalling.
Stem Cell Fate
In the Drosophila germarium, Dpp signals released by the niche promote GSC maintenance by repressing differentiation factors. We found that a complex of Pumillo (Pum) and Nanos (Nos) represses the translation of the translational repressor Brat in GSCs. In the CBs this repression is alleviated, allowing Brat to form a complex with Pum and repress new targets, including Mad, which diminishes the CBs ability to respond to Dpp. Using mathematical modelling we identified bistability of cell fate in the Brat-mediated system, revealing how autoregulation of GSC number can arise from Brat, coupling extracellular Dpp regulation to intracellular interpretation (Harris et al, 2011, Developmental Cell). We also identified the Medea and Schnurri mRNAs as Pum-Brat targets, with post-transcriptional regulation occurring via deadenylation (Newton et al, 2015, Development).
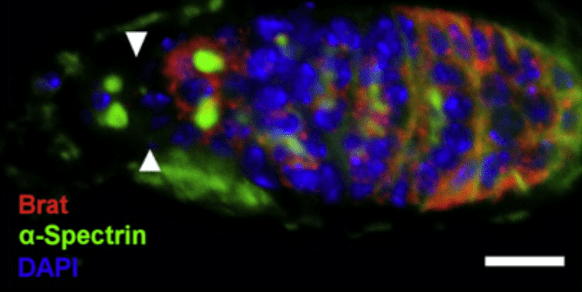
Brat is repressed in the GSCs, indicated by white arrow heads, by a complex of Pum/Nos. In the CBs Brat is expressed and complexes with Pum to repress targets. Scale bar represents 10μm